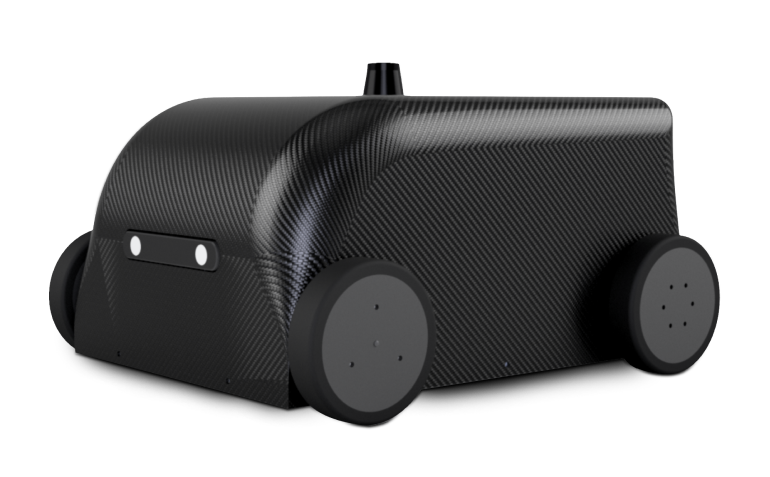
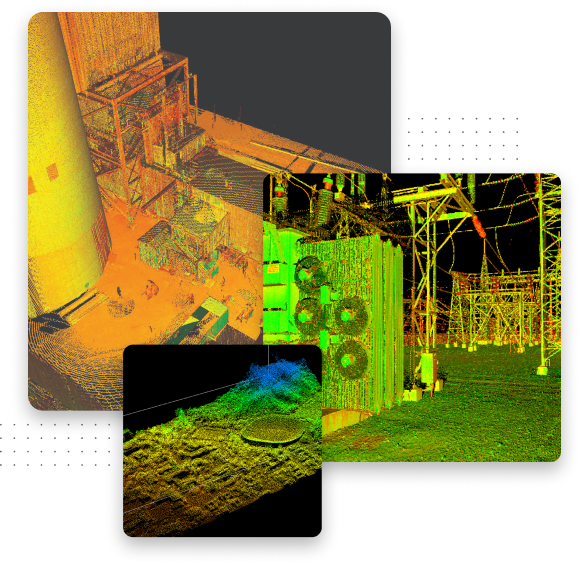
A game-changer for industrial inspection data acquisition and analytics
Quality. Build high-quality, historical datasets and gain deeper insights with automated data acquisition tools for asset management optimisation.
Simplicity. Save time and money with a ready-to-use robotic data acquisition vehicle that doesn’t need a robotics expert to operate.